Modelling unconstrained population growth rates
Let's derive some population growth functions!
This is a rudimentary first-order differential equation which can be solved in a few simple steps.
Solving
While the parameter
library(tidyverse)
trans = matrix(
c(0.00, 0.50, 0.00,
0.00, 0.00, 0.50,
4.00, 0.00, 0.10),
ncol = 3,byrow = T
)
rownames(trans) = c('V1','V2','V3')
colnames(trans) = c('V1','V2','V3')
> trans
V1 V2 V3
V1 0 0.5 0.0
V2 0 0.0 0.5
V3 4 0.0 0.1
Tmax = 200
N = matrix(rep(0, Tmax*3), nrow = 3)
N[1,1] = 1000
for(t in 2:Tmax) N[, t] = N[, t-1] %*% trans
# plot
ages = as.data.frame(t(N))
ages$total = rowSums(ages)
ages$t = 1:Tmax
agesl = gather(ages, age.class, value, -t, - total)
ggplot(agesl) +
geom_area(aes(t, value/total, fill = age.class)) +
ylab('proportion')+
theme_minimal() +
theme(
plot.background = element_rect(fill = rgb(0.2,0.21,0.27)),
text = element_text(colour = 'grey'),
axis.text = element_text(colour = 'grey'),
panel.grid = element_line(colour = 'grey')
)
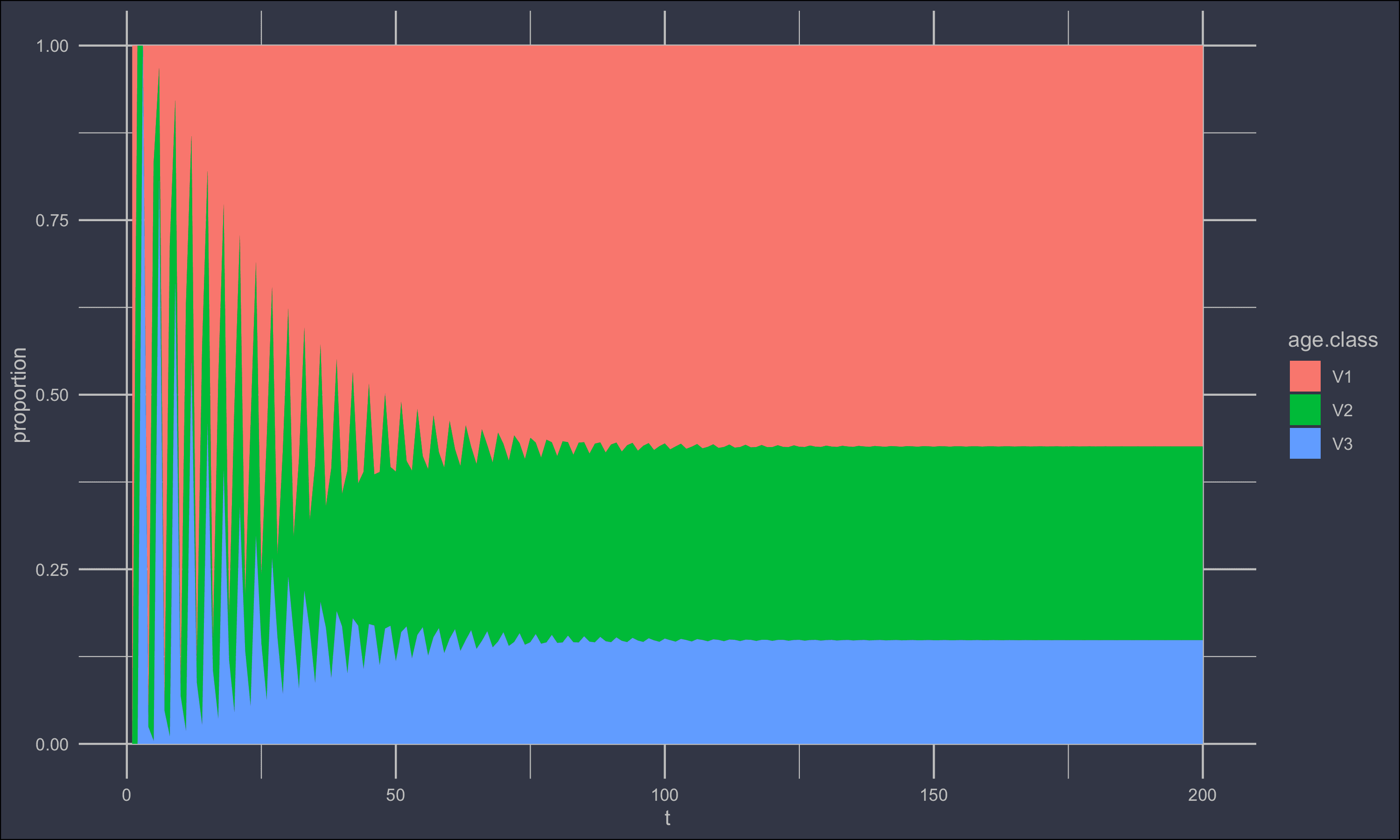
Regressing population vs. time
There are a number of ways that
The most direct way to access
df = data.frame(
t = 0:(Tmax-1),
N = colSums(N)
)
To estimate
lm1 = lm(log(N) ~ t , data = df)
coef(lm1)
> coef(lm1)
(Intercept) t
6.30299077 0.03407767
# plot
df$pred = predict(lm1)
ggplot(df) +
geom_line(aes(t,N), colour = 'red', alpha = 0.5) +
geom_line(aes(t,exp(pred)), colour = 'grey') +
theme_minimal() +
theme(
plot.background = element_rect(fill = rgb(.2,.21,.27)),
text = element_text(colour = 'grey'),
axis.text = element_text(colour = 'grey'),
panel.grid = element_line(colour = 'grey')
) +
scale_y_log10()
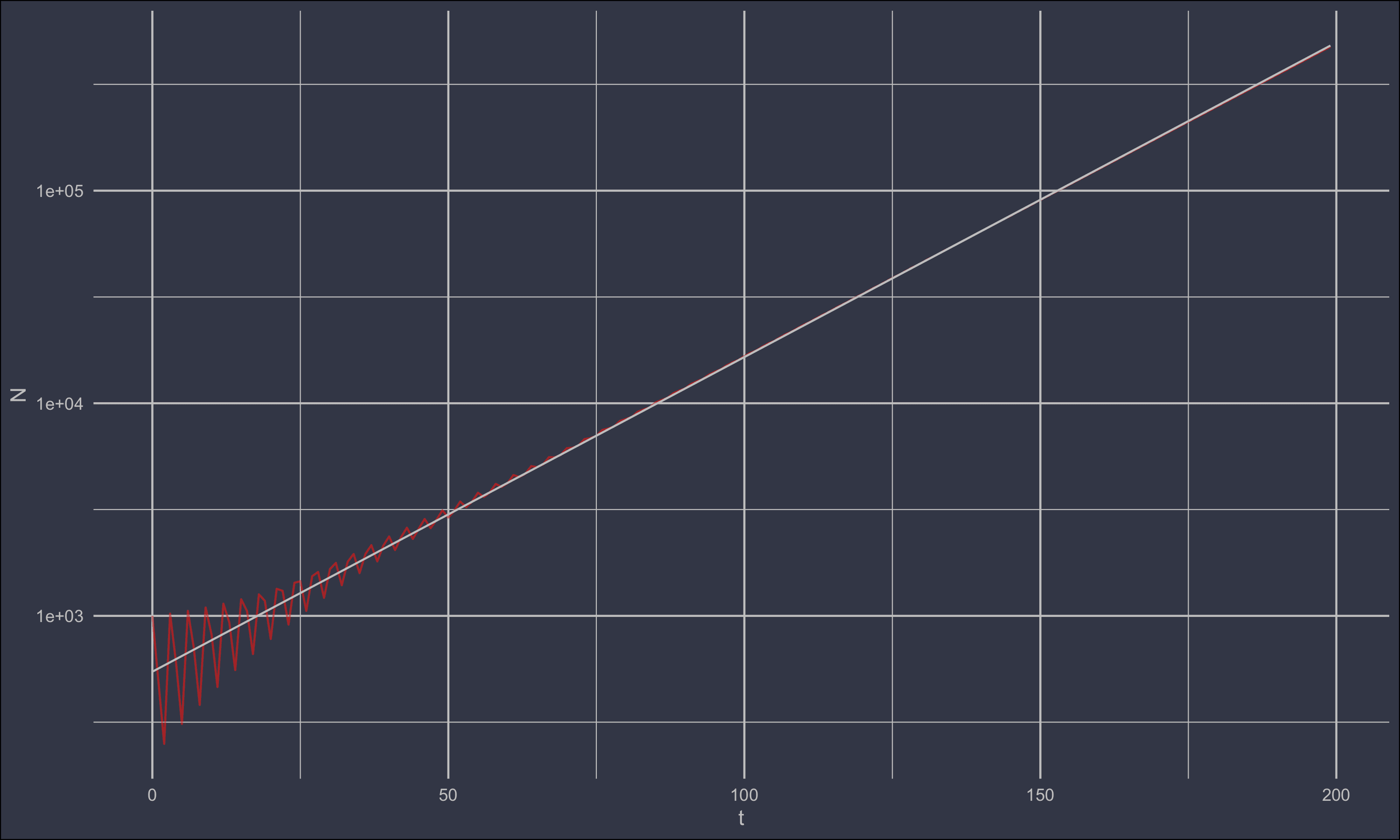
Notice that the recovered intercept exp(coef(lm1)[1]) = r round(exp(coef(lm1)[1])) was actually lower than the real values of 1000. This was due to the age distribution not being in steady state, and the population not growing properly exponential.
Now let's try and run the same matrix model simulation but with the 1000 individuals distributed across age classes at the steady state proportions.
N1_stable = 1000*N[,Tmax]/sum(N[,Tmax])
N = matrix(rep(0, Tmax*3), nrow = 3)
N[,1] = N1_stable
for(t in 2:Tmax) N[, t] = N[, t-1] %*% trans
df = data.frame(
t = 0:(Tmax-1),
N = colSums(N)
)
lm1 = lm(log(N) ~ t , data = df)
> coef(lm2)
(Intercept) t
6.90777973 0.03388836
After exponentiating the intercept we are able to recover the initial population size.
> exp(coef(lm2)[1])
(Intercept)
1000.024
Measuring vital rates and building a life table
Old school population demographers also built life tables for a cohort of individuals followed through life, which include direct measurements of mortality, and reproduction to calculate
Let's use an example published by Carey and Bradley
mite = read_csv('data/AtlanticSpiderMite_CareyBradley1982.csv')
> mite
# A tibble: 26 × 3
x `P(x)` `m(x)`
<dbl> <dbl> <dbl>
1 0 1 0
2 1 1 0
3 2 1 0
4 3 1 0
5 4 1 0
6 5 1 0
7 6 1 0
8 7 1 0.37
9 8 1 3.53
10 9 1 7.43
# … with 16 more rows
# ℹ Use `print(n = ...)` to see more rows
Firstly, we estimate net generational reproduction
# plot
mite$`P(x).m(x)` = mite$`P(x)`*mite$`m(x)`
dw = mite %>% gather(variable, value, -x)
ggplot(dw) +
geom_point(aes(x,value, colour = variable), size = 2) +
geom_line(aes(x,value, colour = variable)) +
theme_minimal() +
theme(
plot.background = element_rect(fill = rgb(.2,.21,.27)),
text = element_text(colour = 'grey'),
axis.text = element_text(colour = 'grey'),
panel.grid = element_line(colour = 'grey')
)
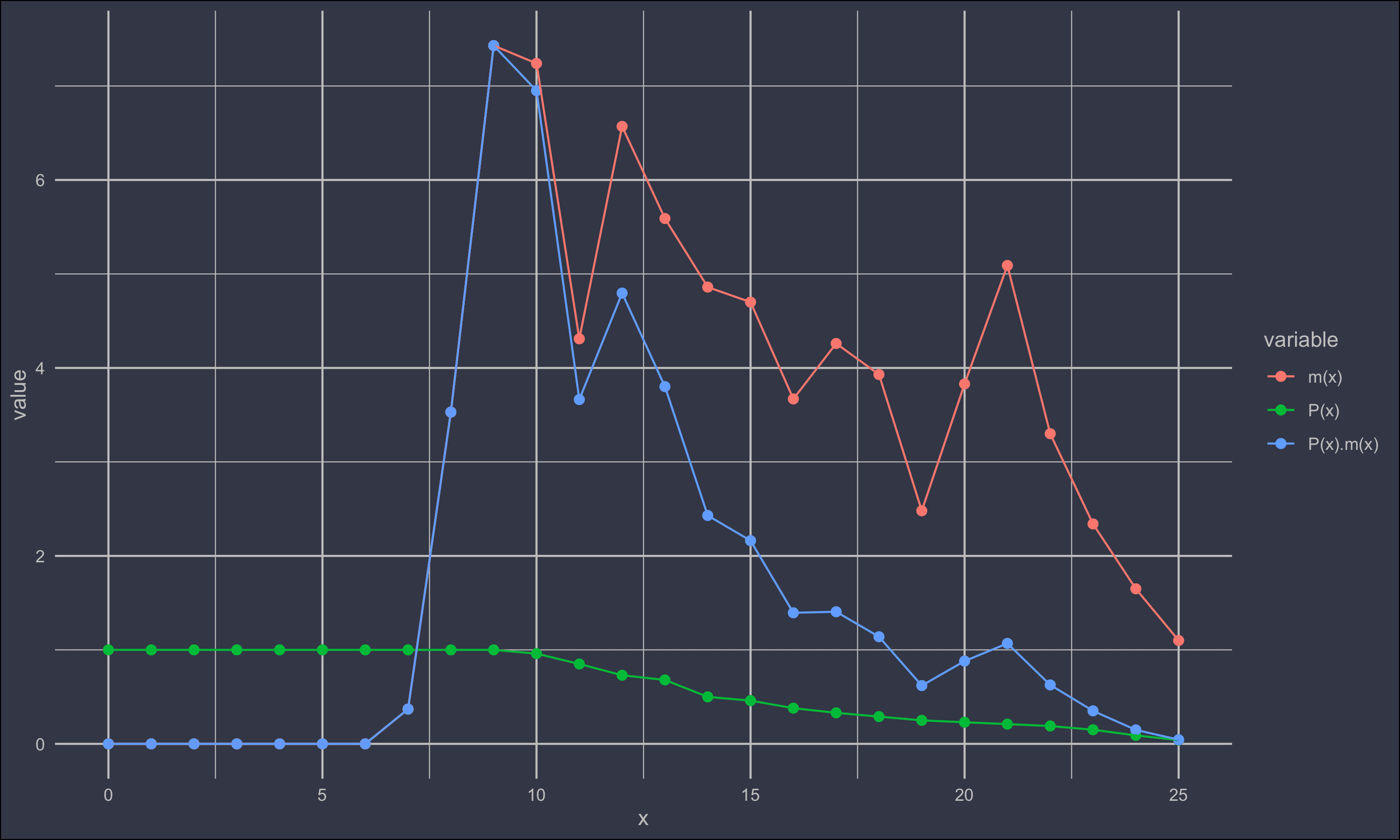
An analytical estimate of r from this data relies on the Lotka equation.
Here we can set
Substituting into the Lotka equation and recognising